 The study by Francavilla and coworkers also used this information to assign each site to one of three clusters (A, B and C) based on its behavior across samples. With the donut visualization we showed the quantitative changes in phosphorylation of each site. How are multiple sites on the same protein shown? Do different donut slices within a protein always show similar changes? 1.5 Pie visualization Select Adj p-value as the column to use for filtering, choose to get to the second dialog, which allows you to select the color palette and fine-tune the color gradient as the default values are fine, simply click Draw to produce the visualization. However, here we will create a simple filter to keep only the phosphorylation sites that are statistically significant at a 1% false discovery rate. In the filter dialog, you can build logical rules that take into account multiple columns in the table. Click the filter button above the network or go to the menu Apps → Omics Visualizer → Filter table. Omics Visualizer allows you to filter the rows in the table before visualizing them on a network. As no changes should be needed for this file, simply click OK to complete the import. The import dialog gives you the option to name the table, shows how the file is interpreted given the current settings, and allows you to change these as needed. Download and open the text file with the data described above. Go to the menu Apps → Omics Visualizer → Import table from file. Start Cytoscape or close the current session from the menu File → Close. An adapted and simplified table with the data from this study is available here. Each protein can have multiple significantly regulated phosphorylation sites, each of which is associated with two log-fold change values (one for each control tissue), a Benjamini–Hochberg adjusted p-value, and a cluster assignment that groups sites with similar behavior across samples. The study used mass spectrometry to compare the phosphoproteome of primary cells derived from epithelial ovarian cancer (EOC) to those of two healthy tissues, namely distal fallopian tube epithelium (FTE) and ovarian surface epithelium (OSE). We will work with phosphoproteomics data from an ovarian cancer study ( Francavilla et al., 2017).  
In this exercise, we will load a data table with proteomics data, filter it, retrieve a STRING network for the proteins, and visualize site-specific information onto the protein network. Similarly, make sure you have enhancedGraphics and stringApp installed before switching back to Cytoscape. Go to the Cytoscape App Store in your web browser and search for Omics Visualizer, select the app and press the Install button to install it. The exercises require you to have certain Cytoscape apps installed. Then start Cytoscape and update the current apps if necessary by checking the App Updates icon in the right-most corner of the manubar. To follow the exercises, please make sure that you have the latest version of Cytoscape installed. visualize time-series data onto networks.visualize site-specific data onto networks. 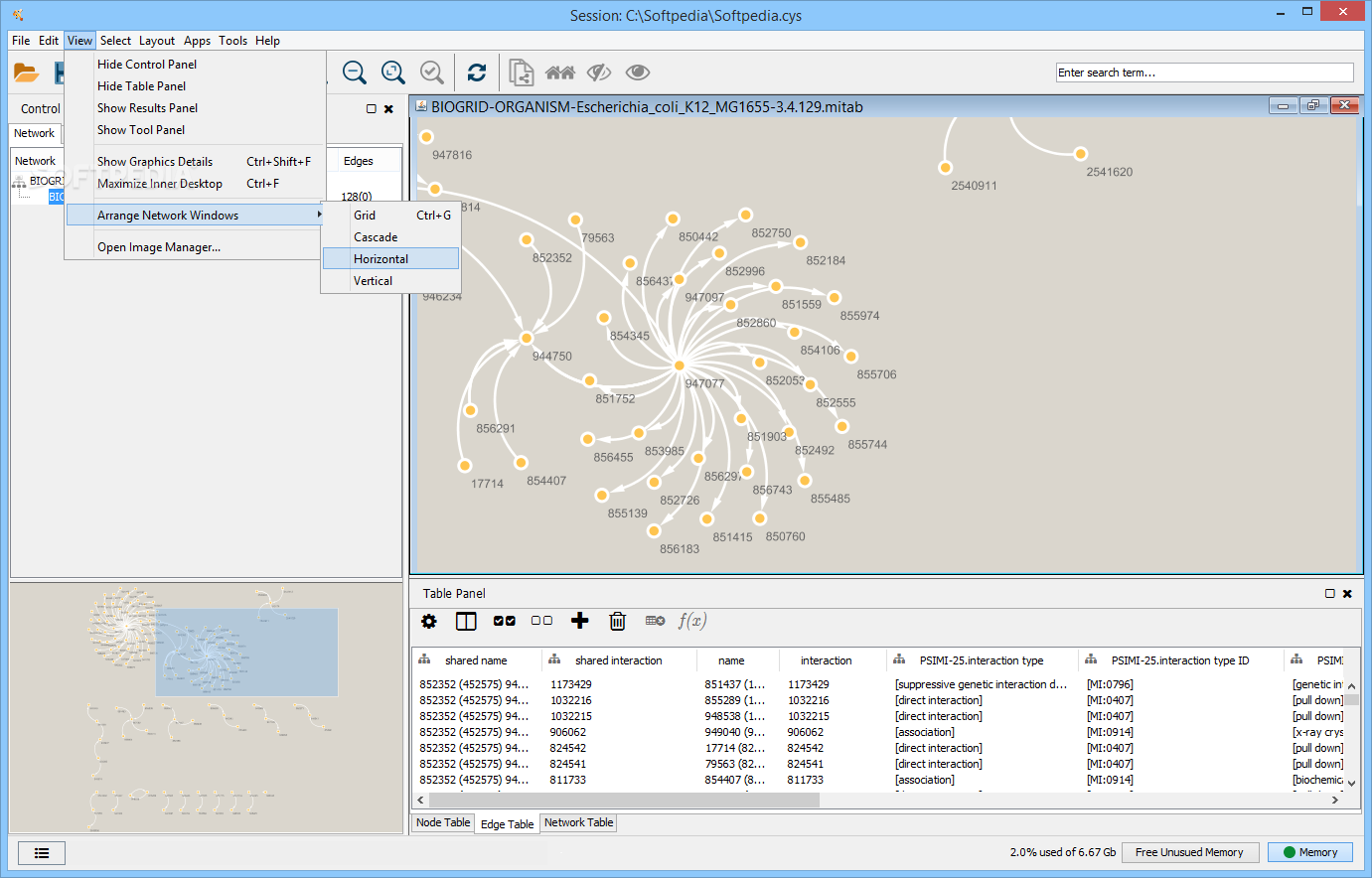
In these exercises, we will use the Omics Visualizer app for Cytoscape to retrieve molecular networks from the STRING database and visualize site-specific information on the nodes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
